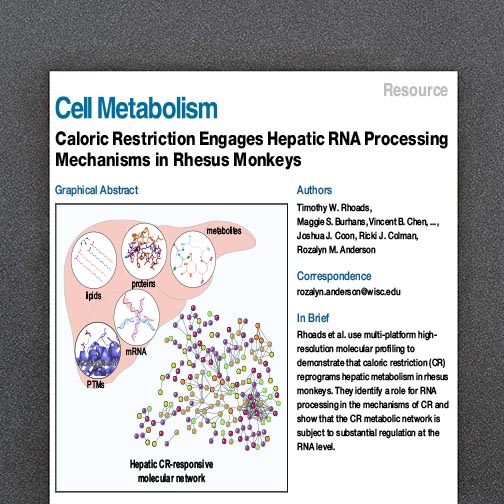
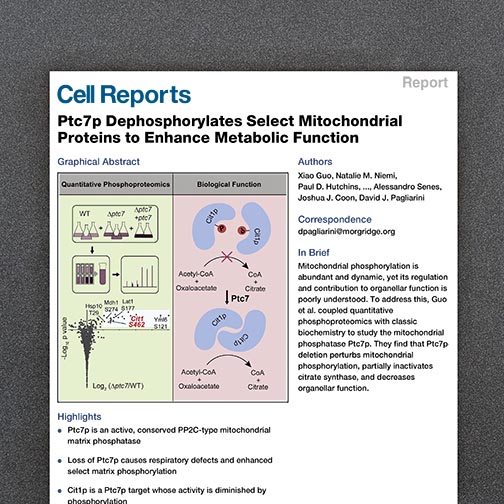
Journal publications provide an important mechanism of communication and dissemination, and we aim to rigorously publish our work as soon as possible, targeting venues having both expert and non-expert audiences. Our overall aim is to promote, advertise, and make available our enabling technologies to the biomedical research community.
Additional resources can also be found at the Biomedical Technology Resources portal, an NIH innovative technology resource.
When appropriate, please acknowledge our support in publications and presentations: “This work was supported in part by the National Institutes of Health grant P41GM108538.”